
Project Goal
Building an atlas of C. elegans worm nuclei in an unsupervised way.
Project Idea:
Consistently matching a large number (100 or more) microscopic worm images with segmented nuclei to each other. The matchings are used in place of ground truth to build the atlas. The project is supported by DFG Projects 98181230 and 53943535 and is run together with Kainmüller Lab in Max Delbrück Center for Molecular Medicine.
Starting Point:
- Building the atlas from partially annotated worm segmentations.
What Was Lacking:
- No fast and accurate enough graph matching (quadratic assignment) solver available — the existing one required ~10 min per worm pair (100×99/2 > 1 month total).
- No fast and accurate enough multi-graph matching solver.
- Supervised pipeline only.

Step 1: Efficient Graph Matching Solver
L. Hutschenreiter, S. Haller, L. Feineis, C. Rother, D. Kainmueller, B. Savchynskyy
Fusion Moves for Graph Matching
ICCV 2021 oral
[pdf]
[project page]
[slides]
[poster]
S. Haller, L. Feineis, L. Hutschenreiter, C. Rother, D. Kainmueller, P. Swoboda, B. Savchynskyy
A Comparative Study of Graph Matching Algorithms in Computer Vision
ECCV 2022
[pdf]
[project page]
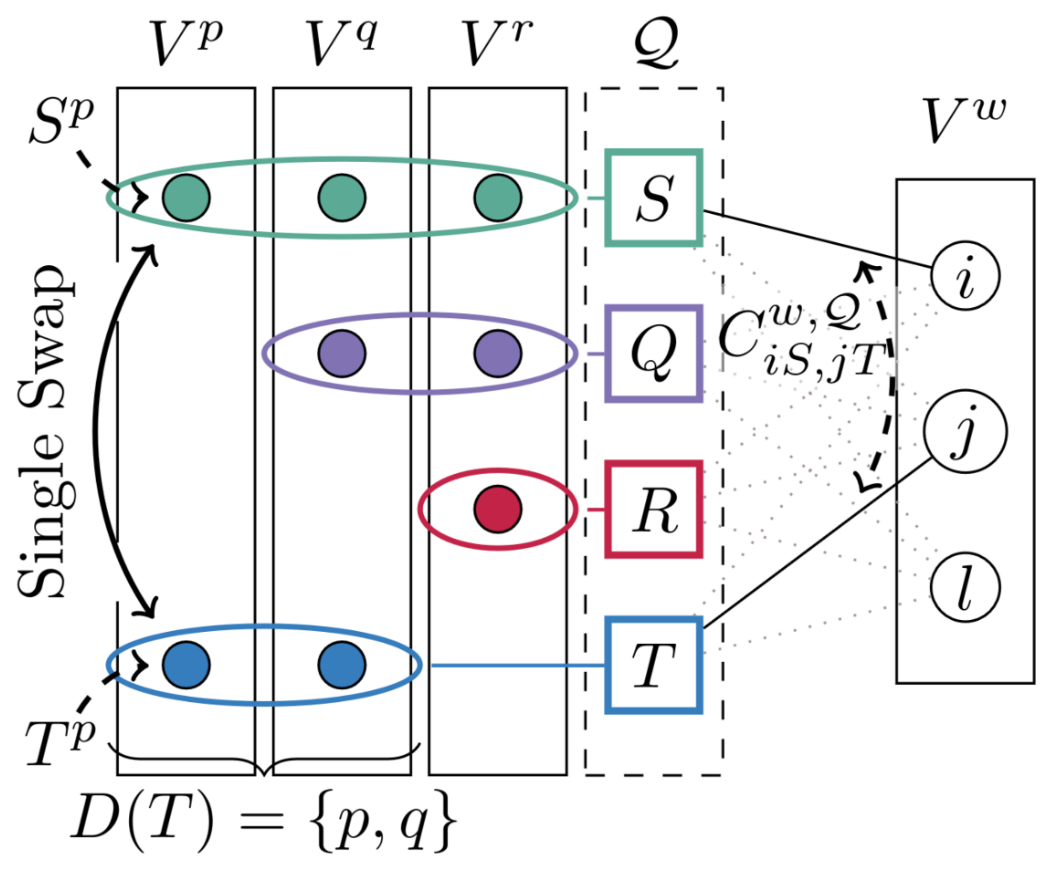
Step 2: Efficient Multi-Graph Matching Solver
Max Kahl, Sebastian Stricker, Lisa Hutschenreiter, Florian Bernard, Carsten Rother, Bogdan Savchynskyy
Towards Optimizing Large-Scale Multi-Graph Matching in Bioimaging
CVPR 2025
[CVPR 2025 version]
[Extended version]
[code]

Step 3: Building the Unsupervised Atlas of C. elegans
Sebastian Stricker, Christoph Karg, Lisa Hutschenreiter, Dagmar Kainmueller, Bogdan Savchynskyy
Cycle-Consistent Multi-Graph Matching for Self-Supervised Annotation of C. elegans
at WACV 2026
[Preprint]
Siddharth Tourani, Carsten Rother, Muhammad Haris Khan, Bogdan Savchynskyy
Discrete Cycle-Consistency Based Unsupervised Deep Graph Matching
AAAI 2024
[AAAI 2024 version]
[extended version]
[code]
[poster]
Current Work
- Integration of Neural Networks into our pipeline to improve modeling power.
- Further improvement of the techniques developed in Steps 1–3.
